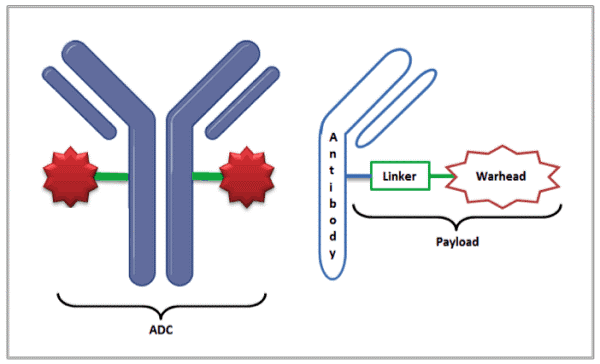
With nearly 10 million deaths in 2020 and despite great progress, cancer remains the leading cause of death worldwide [1]. Antibody Drug Conjugates (ADCs) when first approved in 2000, seemed ideal in that they could deliver the cytotoxic drug (also referred to as the warhead) directly to the tumor site, resulting in reduced off-target damage to healthy tissues. ADCs combine the targeting capability of monoclonal antibodies with the cancer-killing capability of the payload (linker + cytotoxic drug). Today, there are only 11 approved ADCs, but the pipeline for growth in this area is strong, with the global market expected to grow at a compound annual growth rate (CAGR) of 23.7% through 2028 [2].
One way to accelerate the development and approval timeline is to use modeling and simulation, as done successfully by the team at Genentech in the development of Polivy, saving them scarce time and resources [4]. Simcyp’s PBPK (physiologically-based pharmacokinetic) platform provides multiple benefits in development of ADCs viz;
- First-in human dose selection,
- Prediction of drug-drug-interactions (DDI) between the small molecule payload and other co-medications, and
- Understanding the disposition of ADCs in special populations
This allows users not only to optimize their development pipeline, predict the potential hurdles in clinical development but also avoid unnecessary clinical trials. Hence saving significant time & resources which results in additional revenue (with earlier approval).
Here are the Top 10 reasons for using Simcyp PBPK for ADC development:
- The key benefit of using Simcyp PBPK model for ADCs is that our platform models the antibody and the small molecule payload simultaneously.
- Each DAR (Drug Antibody Ratio) is simulated separately, connected via deconjugation. This feature increases the flexibility and simulation accuracy because drug-specific parameters can be individualised for DAR species such as binding to the FcRn receptor or the target. Additionally, other conjugates can be simulated using the ADC model.
- Simcyp also includes a generic tumor model for large and small molecule that allows simulation of the concentration of the DAR species, the naked antibody, and the released payload at the site of action.
- Simcyp provides unique insight into the physiological disposition of ADCs in virtual populations and associated inter-individual variability.
- Simcyp provides users flexibility in the modeling approach i.e., ADCs can be modeled using a minimal PBPK model with a lumped tissue compartment or a whole-body PBPK model that contains 16 different compartments, mechanistically representing different tissues and organs of the human body.
- The full PBPK model for the small molecule payload includes many features of the Simcyp Small Molecule Simulator such as enzyme metabolism, active transport, permeability-limited models, and DDIs.
- The Pharmacokinetic model can be linked to a Pharmacodynamic model to predict the drug effect or any toxicity effect of the payload.
- Simcyp population libraries contain a range of physiological information required to simulate ADCs such as lymph flow and FcRn abundance in each tissue. The included pediatrics population library allows evaluating ADCs disposition.
- Simcyp simulation outputs include information such as individual DAR species, payload release from different sources (deconjugation, clearance, TMDD), tissue penetration etc.
- In addition, sensitivity analysis and parameter estimation are embedded within the Simcyp Simulator and can be used for estimating ADCs drug-specific parameters as well as physiological parameters.
To accelerate your ADCs development pipeline, please contact us!
References
1. WHO. Cancer Key Facts. 2021 [cited 2021 30 September]; Available from: https://www.who.int/news-room/fact-sheets/detail/cancer
2. Research, G.V. Global Antibody Drug Conjugate Market. 2021 [cited 2021 30 Spetember]; Available from: https://www.grandviewresearch.com/industry-analysis/antibody-drug-conjugates-market.
3. Sikorski, S.R.I.Q.R. The Clinical Landscape of Antibody-drug Conjugates. 2014 [cited 2021 30 September]; Available from: https://www.adcreview.com/articles/the-clinical-landscape-of-antibody-drug-conjugates/.
4. Samineni, D., et al., Physiologically Based Pharmacokinetic Model-Informed Drug Development for Polatuzumab Vedotin: Label for Drug-Drug Interactions Without Dedicated Clinical Trials. J Clin Pharmacol, 2020. 60 Suppl 1: p. S120-S131.



