April 1, 2026
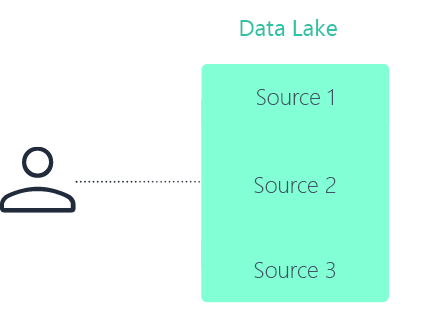
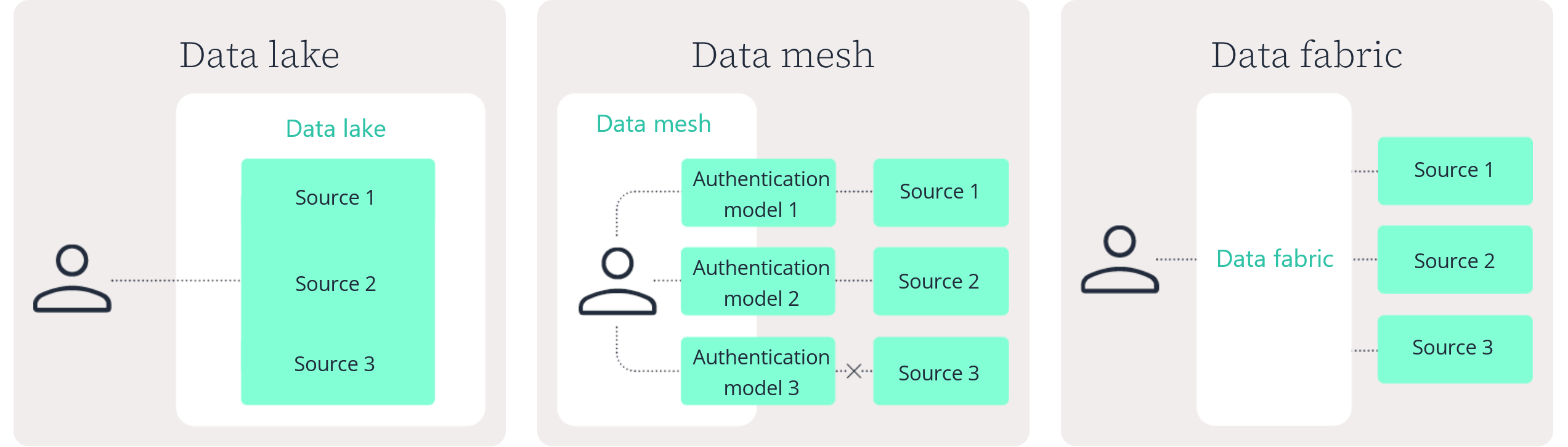
Data lake
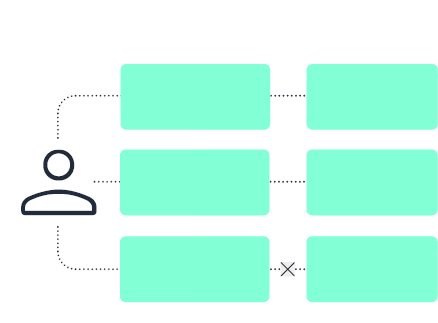
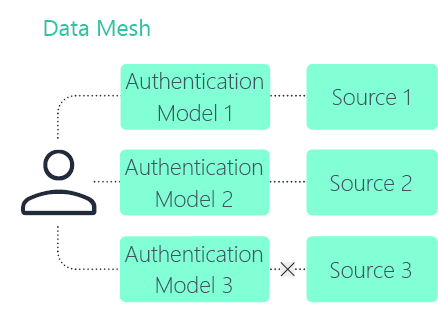
Data mesh
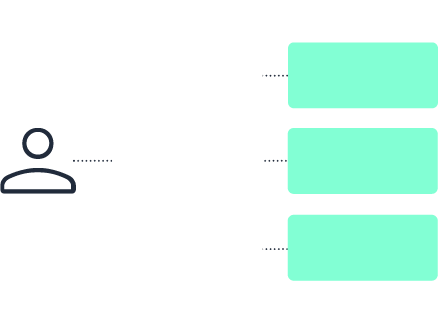
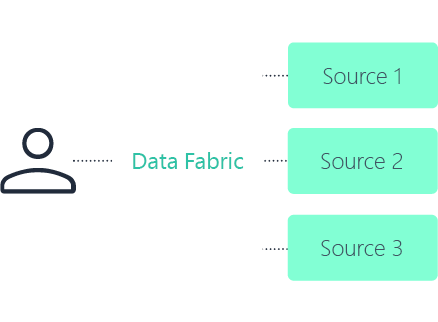
Data fabric
Needle in the Haystack: The Role of Data Fabrics in Successful AI Implementation
Go beyond the concept of data fabrics and see how they are applied in practice. This webinar explores how organizations operationalize data fabrics to support AI, highlighting real challenges, practical approaches, and what actually works.

Author

Sean McGee, MS
Director of Product, CertaraSean McGee is currently the Director of Product at Certara, working within the Certara artificial intelligence (AI) group. Throughout his career, Mr. McGee has supported the strategy and go-to-market motions of various software technologies, including Benchling’s laboratory informatics platform and the AI and molecular modeling and simulation offerings for Dassault Systèmes BIOVIA brand. In his role with Certara, Mr. McGee guides the development of new AI-focused use cases which maximize the benefits of the Certara AI and broader company portfolio.
Mr. McGee completed his Master of Science at the University of Notre Dame exploring the scientific and commercial applications of medical devices designed to aid in the identification of child abuse.
Frequently asked questions about Data Fabrics in Pharma
Is a data fabric the same as a data lake?
No. A data lake centralizes data storage in a single repository. A data fabric in pharma connects data across distributed systems without requiring everything to be moved or duplicated.
A data fabric focuses on unified discovery, governance, and controlled access across environments. A data lake focuses on centralized storage.
When should a pharma organization implement a data fabric?
A data fabric in life sciences is most valuable when data is spread across multiple validated systems, repositories, and document sources that cannot easily be consolidated.
Common triggers include:
- Cross-study evidence search challenges
- Slow clinical or regulatory review workflows
- Heavy reliance on unstructured documents
- The need for AI-ready, permissions-aware retrieval
Organizations often begin with a clinical data fabric (CDF) and expand over time.
Is a clinical data fabric different from a general data fabric?
A clinical data fabric (CDF) is a focused implementation of a broader data fabric in pharma. The architecture principles remain the same, but the connected sources are specific to clinical development.
A CDF typically connects clinical trial systems, safety data, regulatory documents, and related evidence sources while preserving governance and traceability.
Does a data fabric require moving or copying all data?
No. A core principle of a data fabric in life sciences is to connect and index data where it resides. In many cases, systems remain in place for validation, governance, or performance reasons.
The fabric provides a metadata-driven integration layer that enables unified search and access without unnecessary duplication.
How does a data fabric support generative AI in regulated environments?
Generative AI requires reliable context, clear permissions, and traceable outputs. A data fabric in pharma provides governed indexing, metadata, and permissions-aware retrieval.
When combined with retrieval-first workflows and reference-backed outputs, this approach supports faster drafting and analysis while maintaining reviewability and compliance.
Schedule a demo